- Home
- Software
Refine by
Biodiscovery Medical Software
6 software items found
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
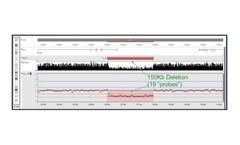
As described in Chaubey et al., Journal of Molecular Diagnostics, vol. 22, No. 6 June 2020, they used 10x WGS and validated that the NxClinical algorithm detected all CNVs and AOH that were found by high-resolution SNP arrays. Figure 2. shows a small exonic deletion detected using 10x WGS with the MSR algorithm. With higher depth NGS, smaller CNVs can be detected and integrated with sequence ...
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
The most comprehensive and up-to-date solution for cytogenetics and molecular genetics in one system for analysis and interpretation of all genomic variants from microarray and NGS ...
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
The most recent version of NxClinical, the leading copy number variation (CNV) analysis software for clinical cytogenetics and molecular labs, now includes an integrated ACMG guidelines scoreboard ...
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
Arming researchers with advanced genomic data analysis and visualization tools. Easy-to-use software provides cutting-edge statistical tools to advance scientific ...
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
BioDiscovery has developed two algorithms for the detection of CNV and AOH events from NGS using its decades-long expertise in the area. One algorithm, the “Self-reference” algorithm, can be used for all WGS data regardless of sequencing depth. The second is the “Multi-Scale Reference” (MSR) algorithm that is applicable to all NGS data (WGS, WES, and Gene panels). The MSR ...
by:BioDiscovery, A Bionano Genomics Company based inEl Segundo, CALIFORNIA (USA)
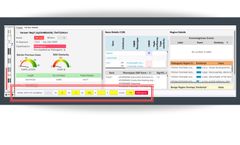
The MSR algorithm can be applied to any gene panel from single gene (e.g. DMD test) to large panels having thousands of gene. The image below is from the Illumina TruSightTM Oncology 500 (TSO500) panel showing a somatic cancer profile. The cytogenetic complexity of the tumor sample is clearly evident with a large copy number gain of 8p and loss of a large section of 13q. Aberrations associated ...