

SOPHiA - Version DDM -Coud-Based Software Platform for Rare Diseases
Next-generation sequencing (NGS) of the human exome has the potential to identify pathogenic variants responsible for complex phenotypes associated with rare diseases.
advanced analytics and dedicated features
Quickly and accurately detect variants associated with rare diseases with the advanced analytical capabilities and dedicated features of the SOPHiA DDM Platform complemented by Alamut™ Visual Plus. Streamline your exome data-driven research today.
Mutations in mitochondrial DNA (mtDNA) cause a diverse range of diseases affecting 1 in 5000 people,2 a substantial portion of the rare disease community. NGS of the mitochondrial genome, alongside the exome, is therefore valuable in rare disease research. In a single workflow, the SOPHiA DDM Platform analyzes variants in mtDNA alongside SNVs, Indels, and CNVs in both coding and clinically-relevant non-coding regions of nuclear DNA. The platform’s sophisticated algorithms address the unique challenges associated with the mitochondrial genome such as the variable amount of mtDNA and heteroplasmy, to identify causative variants.
Exome sequencing can detect tens of thousands of variants, and determining which variants are associated with a rare disease phenotype can be a daunting task. Virtual Panels, Familial Variant Analysis, and Cascading Filters are user-friendly, time-saving features in the SOPHiA DDM Platform that can prioritize variants for further analysis.
Virtual Panels: Integrated databases such as HPO automatically select genes associated with the suspected disease.
Familial Variant Analysis: Variant haplotypes are displayed in DDM with the option to filter according to inheritance mode.
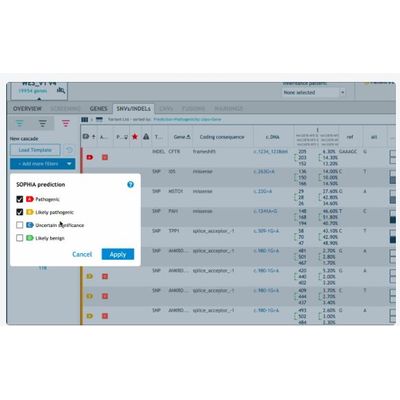
Cascading Filters: Custom strategies can be set up in a few clicks using filters such as ACMG score, ClinVar pathogenicity, and SOPHiA GENETICS prediction.
When you are ready to focus on your variants of interest, Alamut™ Visual Plus allows you to connect the dots by exploring them on a genomic scale in a comprehensive, full genome browser supported by world-renowned curated databases, guidelines, and splicing predictors. The intuitive and user-friendly interface allows visualization of variants in GRCh37/38 assemblies and the mitochondrial genome, while conveniently displaying flanking regions and overlapping genes.
